Disclaimer: I am not an expert in international relations or military strategy, which is fine. In democracies, it’s normal and correct for ordinary citizens to have opinions on important world issues, and demands that they not do so are ahistorical and dangerous. Still, take anything I say with a grain of salt.
1: This isn’t “history restarting” . . . yet
Whatever Francis Fukuyama meant by “the end of history”, it probably wasn’t “nothing will ever happen”.
But that’s how it’s been interpreted, so fine. Maybe nothing will ever happen. I don’t think the Ukraine War is necessarily a counterexample. Fukuyama wrote in 1992, so he knew that eg the Gulf War could happen. Is this conflict bigger than the Gulf War?
I don’t think Ukraine proves that “history has restarted” or “the Pax Americana was a paper tiger” or anything of the sort. These kinds of local conflicts were always allowed. Just ask an Iraqi. Or a Chechen, or an Afghan, or a Syrian, or a Bosnian, or a Crimean, or a Tigrayan or go back and ask the Iraqi a second time.
But the vast majority of people reading this have probably never been personally affected by a war and might not even know anyone who has been. And a billion Chinese, and almost a billion Indians, and almost everyone in South America, and a lot of other people, can say the same.
Outside of lulls in history and Pax Somewheres, one nation invading another is met with indifference. Russia’s invasion of Ukraine is being met with internal protests, global condemnation, and crippling economic sanctions. This is what it looks like when a civilization that’s got strong and well-functioning norms against aggressive wars encounters one and launches an immune response.
2: If the Pax Americana is dead, we need to try something different; but if it’s still alive, we should stick with what works.
The Pax Americana playbook for international norm violations is: the US slaps sanctions on the offender. The EU expresses “concern”. The UN proposes a resolution condemning it, which gets vetoed by whichever Security Council member is most complicit. And the CIA secretly gives Stinger missiles to everyone involved.
Lots of people have lost faith in the Pax Americana, which would mean we need something other than the playbook. These people tend either towards extreme isolationism, where sanctions are an aggressive act and even expressions of deep concern are violations of national sovereignty. Or towards extreme bellicosity, where we’re cowards unless someone puts boots on the ground and start shooting the perpetrator directly.
But if the Pax Americana still holds, then the playbook is still the right call.
3: A strong response right now isn’t just about Ukraine, it’s also about the next time.
Everyone can sanction Russia as much as they want, and it can win anyway. Putin is in too deep to extricate himself easily; it’s become a matter of honor, of “not being seen to be weak”. The point isn’t to save Ukraine, it’s to establish expectations for next time. This is about Taiwan, Georgia, Iran, and all the other places that great powers want to invade but don’t.
The next time a big country wants to invade a little one, we want it to remember how much misery everyone inflicted on Russia for the Ukraine conflict and think “no thank you”. That involves inflicting lots of misery on Russia right now, whether or not it wins the current war.
This is true not just for the West considering sanctions, but for Ukrainians considering how hard to fight. Commentators have drawn connections between the Taliban easily ousting the US-backed Afghan government, and Putin expecting an easy victory in Ukraine. Maybe that’s why he took the chance. But the heroic Ukrainian resistance will set the opposite example. Next time someone considers an invasion, they’ll expect such high costs it won’t be worth it. In this sense, the Ukrainians are sacrificing not just for their countrymen, but for the world and for peace itself.
4: International norms may be annoying, but they’re all that stands between us and nuclear war, so we had better respect them
If you only get one thing from this essay, let it be: unless you know something I don’t, establishing a no-fly zone over Ukraine might be the worst decision in history. It would be a good way to get everyone in the world killed.
The “usual playbook” can seem half-hearted and faintly ridiculous. “We’re Not Participating!!!” we insist, as we provide guns and missiles to the people who are. It feels like a bunch of arbitrary lines where we act with bluster and bellicosity on one side, then shrink like fainting violets away from the other. But those arbitrary lines are what save us from global annihilation.
Any sane person wants to avoid nuclear war. But this makes it easy to exploit sane people. If Russia said “Please give us the Aleutian Islands, or we will nuke you”, what should the US do? They can threaten mutually assured destruction, but if Russia says “Yes, we have received your threat, we stick to our demand, give us the Aleutians or the nukes start flying”, then what?
No sane person thinks it’s worth risking nuclear war just to protect something as minor as the Aleutian Islands. But then the US gives Russia the Aleutians, and next year they ask for all of Alaska. And even Alaska isn’t really worth risking nuclear war over, so you give it to them, and then the next year…
So people who don’t want to be exploited occasionally set lines in the sand, where they refuse to make trivial concessions even to prevent global apocalypse. This is good, insofar as it prevents them from being exploited, but bad, insofar as sometimes it causes global apocalypse. So far the solution everyone has settled on are lots of very finicky rules about which lines you’re allowed to draw and which ones you aren’t.
If there was ever a point at which two nuclear powers disagreed about who was in the wrong, one of them could threaten nuclear war to get that wrong redressed, the other could say they had drawn a line in the sand there to prevent being exploited, and then they’d have to either back down (difficult, humiliating) or start a nuclear war (unpleasant, fatal). So there are a lot of diplomats who have put a lot of effort into establishing international norms on which things are wrong and which things aren’t, so that nobody crosses anyone else’s lines by accident.
This system isn’t perfect. Nuclear powers disagree on lots of things. But they usually disagree in a bounded way, where they accuse each other of non-mortal sins and claim the right to non-nuclear responses. Russia crossed a line by invading Ukraine, in a way that gives Russia’s enemies the right to certain kinds of retaliation - arming Ukraine, imposing sanctions, etc. Russia will grumble about this, but it knows it would be in the wrong if it threatened a nuclear response - it would be violating the West’s lines in the sand, the West would have to call its bluff, and it would have to either go ahead with apocalypse or back down in humiliation.
I am not an international relations expert. But every international relations expert whose commentary I have read claims that the extent of Russia’s recent infraction does not give the West the right to declare a no-fly zone in Ukraine. The no-fly zone would be an extreme escalation that would, under international norms, allow Russia to threaten World War III if we didn’t back down. Then we would either have to back down, humiliated, or start World War III. In a situation like that, I pray we would have the courage to back down humiliated. But I would prefer not to test our leaders’ courage in this particular way.
Also, the last time this happened, in ‘62, it was the Russians who agreed to back down to prevent nuclear war. We owe them one, so this time it’s on us.
5: Really, I can’t emphasize this enough, a no-fly zone means shooting down Russian planes.
America does not actually have a way to prevent people from flying. A no-fly zone means that if they do fly, you shoot them down. It would be more reasonable to call this a “shoot-down-anything-that-flies zone”, but at some point some Pentagon official must have wanted to sell it to the public really hard and come up with an innocuous-sounding name for it.
If America actually shoots down Russian planes, there is a decent chance it causes World War III. At the very least, our strategy for preventing World War III would be “shoot them, hope really hard that they don’t shoot back”, WHICH IS NOT A GOOD STRATEGY.
But isn’t it possible that the US could declare the no-fly zone, and the Russians (who also don’t want World War III) would agree not to fly, rather than cause global annihilation?
I think this is where the lines-in-the-sand come in again. Imagine Russia declared a “no-sanctions zone” across the entire world, where if any corporation stopped doing business with them, they would bomb that corporation’s headquarters (even if the corporation was headquartered in eg the US). While this might give some corporations pause, a lot of Americans would feel honor-bound not to comply - it would be “giving into terrorism”.
The line between common-sense “don’t provoke a nuclear power” and “if we went along with this, it would be giving into terrorism” is set by international law, diplomatic norms, and various fuzzy rules of war. They say that some things are allowed, and other things are bullying and if someone threatens them you need to call their bluff. The silly “no-sanctions zone” idea would be the latter. And so would a no-fly zone.
Putin’s already proven a little irrational. He’s done good work establishing himself as the sort of person who calls all bluffs that it’s in his interest to call. So stop trying to put him in a position where sticking to his usual habits would cause World War III.
I also feel this way about letting Ukrainian jets use NATO bases, and anything else that diverges from the usual rules of noncombatants.
6: Huh, I guess we’re still capable of jingoism
One story you could tell - one story I think Putin was telling himself - goes something like: the West is pathetic and divided. The Western-backed Afghan government fell to the Taliban in a few weeks. That’s because the Westernized Afghans were the kind of people who cared about trigger warnings and misgendering, and the Taliban was Traditional Masculine Warrior Types. The Taliban could say “kill the infidels!” and the Westerners would argue over whether considering the Taliban an “enemy” was racist.
The past few years have seen some of the most powerful players in the Western world, like the big tech companies, refuse to help their own military because they think it’s evil. They’ve seen American conservatives say nice things about enemy dictators because at least they’re not American liberals, and American liberals start treating guns like some kind of eldritch artifact that makes anyone who touches them or associates with them inherently polluted.
So a totally reasonable story would be that the West has become psychologically unsuited for war. Ukrainians would be unable to fight (at least successfully), and Westerners would be too complacent to unite behind Ukraine, especially if it meant higher gas prices or whatever.
(before the war, I saw people on both sides overestimating the relevance of the Azov Battalion, ie the neo-fascists, on the grounds that only neo-fascists would have enough traditional values left to put up a real fight)
That story has fallen apart in two ways. First, the valor of the Ukrainian people. I’m sure there will be debate over whether this is because Ukrainians aren’t as Westernized as Americans, or whether Westernization is more compatible with martial valor than previously expected. But it sure is a data point.
Second, the - let’s call it jingoism - of the broader West. I want to be clear here: so far, Westerners have not actually displayed any martial valor. They’ve mostly displayed the ability to be really pro-Ukraine on Reddit. Still, they sure have been really pro-Ukraine on Reddit. All the people who used to post cringeworthy comments about “Drumpf” are posting cringeworthy comments about “Putler”. I wouldn’t believe it if I hadn’t seen it with my own eyes.
Even so, I think this demonstrates an ability to unite against a foreign enemy beyond what me-a-month-ago would have expected. I think if we ever get in a really important war, we will do just fine on the home front.
Of course, jingoism is bad. People are going crazy, trying to take out their frustrations on individual Russians, or agitating for nuclear war, or otherwise embarrassing themselves.
All of this is terrible. But I was so concerned we were perma-stuck at the opposite extreme that it’s almost refreshing to see us fail in this particular way.
7: The Obligatory Acknowledgment That We Are Also Bad
America has invaded a lot of countries, even within my lifetime.
Sometimes its reasoning was noble: preventing genocide in Kosovo. Sometimes it was at least understandable: get vengeance for 9-11. Other times it was almost incomprehensible: we’ll debate what happened with Iraq II forever.
Part of me wants to say we’re different from the Russians - at least we haven’t launched a war of annexation in a while. Usually we have the decency to skulk around funding rebel groups and opposition parties instead of launching full scale invasions. The rare exceptions tend to be genuinely bad dudes - however unjust the Iraq War was, nobody wants to defend Saddam.
But there’s a failure mode where every villain can come up with at least one rule they followed which the other villains didn’t, then guiltlessly condemn the other villains for their villainy. Putin says that invading Ukraine is okay, because they’re Nazis; maybe he even believes it.
There’s a constant tension between axiology/consequentialism/Inside View morality and law/deontology/Outside View morality. The former says “the Good is indescribably complex, but you can usually recognize it when you see it; follow the Good and ignore the heuristics”. The latter says “bargain with other people until you find bright-line rules you can all agree on, then follow them.”
So, how bright-line rule is “never invade another country”? If the other country had a universally-hated dictator who was genociding millions of people, and it would be easy to invade them, and God Himself came down and assured you that nothing would go wrong - do you wash your hands of it and say “nope, there’s a bright line against ever invading another country, that’s Morally Wrong”?
What if, whenever you admit an exception to the bright line, you know that tyrants and aggressors will exploit it forever? “Fine, you can invade if the country is literally the Nazis, committing the literal Holocaust” - and then Putin says Ukraine is run by Nazis and genociding its people.
I have no good solution to this problem, but I admit that America’s standing to make the moral case against invading Ukraine is weaker than if it had shown the slightest ability to refrain from invading places it wanted to invade.
8: The Obligatory Acknowledgment That We Are Also Bad (2)
What to make of the claim that the West provoked this war by expanding NATO / refusing to rule out admitting Ukraine?
This is one of those times you have to be really careful with causal vs. moral language - in a purely historical sense, did the West cause the war by expanding NATO? And, as a separate question, does this make the West blameworthy?
(in case the distinction isn’t clear, a woman wearing skimpy clothing might be causally responsible for her being sexually harassed, but doesn’t make her blameworthy)
I found Cuban Missile Crisis analogies helpful here: the US also gets nervous when enemy powers are right on its doorstep. So it’s not crazy for Russia to be worried. Still, Putin also uses a lot of “Ukraine is a fundamentally illegitimate country whose very existence is an affront to Russia” rhetoric. Seems like he has a beef besides potential NATO membership (which everyone agreed wasn’t really going to happen).
But also, Russia keeps trying to turn nearby countries into puppet states, sometimes propping up really abhorrent dictators (eg Lukashenko) to do that. They already invaded Ukraine once, took some territory, and propped up some separatist movements. If Ukraine avoided requesting Western connections and military help, or the West avoided providing it, I think “Ukraine becomes Belarus 2” is more likely than “everything is great and war is averted with zero problems”.
Is it wrong for the West to support Ukraine in its efforts not to become Belarus 2? In terms of the lines-in-the-sand and vague-rules-of-international-diplomacy that prevent nuclear war, I think not really. Is it imprudent? It’s a risk, but at least it was taken in the defense of real principles, which is better than most of the imprudent things we do.
9: Peace is still the goal
Putting all of this together: Western countries have three conflicting goals here. First, avoiding nuclear war. Second, making this such a miserable experience for Russia that nobody tries anything like it again. Third, helping the people of Ukraine (and Russia) escape with as little death and suffering as possible.
In this spirit, I hope they encourage Ukraine to consider Russia’s recent peace offer.
As far as I understand it, the offer is: Ukraine declares neutrality, and recognizes Crimea as Russian and Donetsk/Luhansk as independent. Russia gives up and goes home.
These are concessions in name only. Russia already has de facto control of Crimea, Donetsk, and Luhansk, and has for years (I’m assuming Putin means the areas he already controls; if he means “Donetsk” and “Luhansk” in a broader sense, that’s a harder sell). Ukraine ceding them does nothing except take away Russia’s casus belli for future wars.
My understanding is that Russia operationalizes neutrality as “don’t join NATO or EU”. But NATO has shown no signs of being willing to accept Ukraine as a member anyway. EU seems sort of willing, but is infamous about dragging membership negotiations out for years or decades, and requires potential members to get their acts together to a degree that Ukraine might never accomplish. EU has previously allowed members to join its economic community without joining the EU proper, and this would probably provide most of the relevant benefits to Ukraine without angering Russia. Ukraine was not in either of these organizations before the war, and not being in them afterwards changes nothing.
(Neutrality does prevent them from gaining useful allies for a future war, but the only people who might declare war on them in the future are Russia, and they’ve already made it clear they’re a tough target. Some analysts say Putin attacked now because, given the rate at which Ukraine’s military is improving, he thought this would be his last chance. Given the popularity boost and boost in foreign interest this war will give them, they’ll only improve faster from here, so I think they should expect to be able to stand on their own in the future.)
These were most of Putin’s demands before the war, so one could argue that, if they’re a good idea now, they would have been a good idea then, and Ukraine should have agreed and prevented bloodshed. This might be true, but I don’t think it’s necessarily so. A big part of diplomacy is maintaining your honor and your reputation for not caving in to threats. If Russia had gotten everything it wanted from Ukraine with no effort, it would have legitimized using threats of war as a negotiating tactic and made it harder for Taiwan/Georgia/Iran etc in the future. This is easy for me to say on the other side of the world, not losing any friends or relatives, but it is potentially worth standing up for yourself, even to the point of war, in order to maintain the illegitimacy of such threats.
Now the situation is different. Russia has miscalculated, they know they’ve miscalculated, and the best ending for everyone is for them to leave in a way that sort of preserves what’s left of their honor - one that doesn’t humiliate them any more than they’re humiliated already. Giving Russia everything they wanted before the war lets Putin play it as a victory back home, saves the Ukrainian people, and defuses the chance of World War III. It might cost a small amount of honor, but the Ukrainians are rolling in honor right now. They have so much honor they don’t know what to do with it all. They can pay a little to make Russia go away, and still have enough left over to act as a deterrent in the future.
10: Links
a. Metaculus Alerts is a Twitter bot that alerts you when a Metaculus prediction on the Ukraine war has changed drastically in a short time. For example, “the chance of Russia taking Kiev by April has decreased 10% in the past 24 hours”. I find this a good substitute to refreshing the news every minute to see if something interesting has happened.
b. The origin of “Molotov cocktail”:
c. One of the Ukrainian cities on the front lines is named New York.
d. Reddit has quarantined their r/russia subreddit, which I think is a cowardly and outrageous act of censorship. But you can still see it if you have a verified email, and I find it an interesting window into the Russian perspective on the conflict.
e. Former oligarch Petro Poroshenko is Ukraine’s unpopular ex-president, recently placed under something like house arrest pending a corruption trial. He’s since gotten an Kalishnikov rifle and is patrolling the streets of Kiev against Russian invaders.
f. The Reply Of The Zaporizhian Cossacks is a famous historical insult sent by Ukrainian cossacks to the Turkish sultan (it’s worth clicking the links for the full text, content warning obscenity). It got made into a famous painting, and:
g. Maybe Russian propaganda, but still pretty funny:
h. Re: “the West is turning cancellation into a weapon of war”
i. Elon Musk sends Starlink terminals to Ukraine to ensure continued Internet, although there are worries that Russia can trace the signal. Pic related:
j. In Greek mythology, Snake Island, where Ukrainian soldiers famously defied a Russian warship, is the final resting place of Achilles, who sometimes appears to residents.
k. Metaculus thinks Russia might soon close its borders. It might be helpful to talk to Russians you know about getting out of Russia if they can, before things get worse. See also Letter: Russians Are Welcome In America - though I don’t know what the visa situation is like now and it might be terrible.
l. Servant Of The People is a 2015 Ukrainian comedy TV series about a poor teacher who implausibly gets elected President of Ukraine and has to clean up its corrupt politics. It went down in history when the star, Volodymyr Zelenskyy, got elected President of Ukraine in real life, apparently on the strength of his performance. The entire series is available for free on YouTube with English subtitles (though after a few episodes they disappear, and you have to use the Russian subtitles and then auto-translate them into English). I’m a few episodes in and it’s really good, which I guess I should have predicted given the consequences.
m. The EA Forum and Kelsey Piper have discussions on how best to help Ukrainians (this is still not the most efficient way to spend charitable donations - but it’s human to care about things other than efficiency). Ideas range from Polish Humanitarian Action (to help Ukrainian refugees in Poland) to Meduza (opposition Russian news source, apparently still sort of holding on) to direct donations to Ukraine’s Ministry of Health or Ministry of Defence.












 Oliver Traldi @olivertraldi
Oliver Traldi @olivertraldi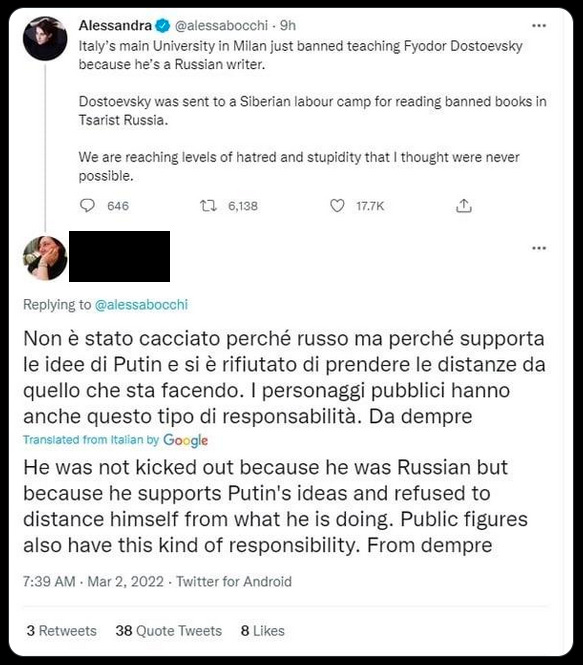
 Aella @Aella_Girl
Aella @Aella_Girl
 Chris Strider @stridinstrider
Chris Strider @stridinstrider

 Anatoly Karlin (Z,Z) @akarlin0
Anatoly Karlin (Z,Z) @akarlin0 The Washington Post @washingtonpost
The Washington Post @washingtonpost International Cat Federation bans Russian cats from competitionsThe Fédération Internationale Féline is among the latest groups to isolate Russia after its invasion of Ukraine.wapo.st
International Cat Federation bans Russian cats from competitionsThe Fédération Internationale Féline is among the latest groups to isolate Russia after its invasion of Ukraine.wapo.st